
|
You can search for RNAs typing in the text area the molecule of interest (e.g. hsa-miR-21) or you can browse the database. When you start typing the molecule, the system will suggest an entry in our database: |
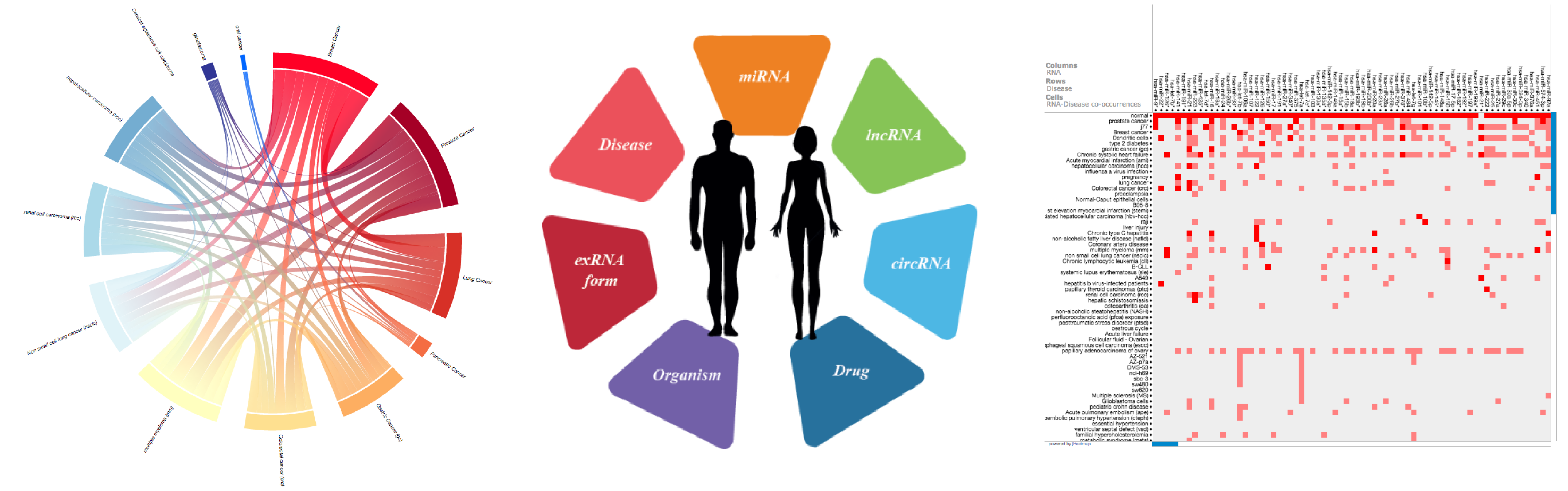
miRandola is a comprehensive manually curated classification of different extracellular circulating non-coding RNA types. The first version of the miRandola database was published in 2012. It contained 89 papers and only miRNA data from humans. Now, we have updated the database with 314 papers! We are starting to add more RNA molecules such as lncRNAs and circRNAs.
Extracellular RNAs (e.g. microRNAs) in serum, plasma, saliva, urine and other body fluids have recently been shown to be associated with various pathological conditions including cancer. RNAs circulate in the bloodstream in a highly stable form, thus they may be used as blood-based biomarkers for many diseases.
The miRandola database 2017 includes (last update February 2017):
The first version of the miRandola database has been released in April 27, 2012. It appeared for the first time at the annual meeting of the Bioinformatics Italian Society 2012 (Catania, Italy). After less than 5 years, miRandola received 100 citations (source: Google Scholar citations, January 2017). Awards: Francesco Russo received the Best Poster and Travel Fellowship Awards presenting miRandola at The Ninth International Biocuration Conference (2016), Geneva, Switzerland (follow the link to the F1000Research channel https://f1000research.com/posters/5-724).
Reference